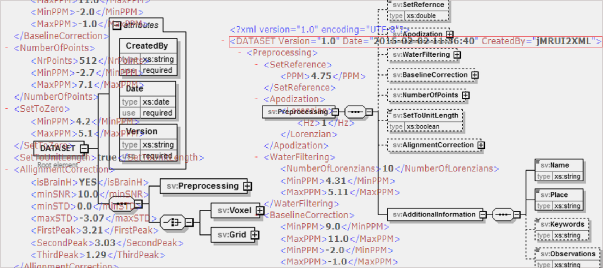
Take note of the incoming virtual meeting “SpectrIm – a tool for the combined analysis of MR Spectroscopy and Imaging” presented by Dr. Johannes Slotboom and Nuno Pedrosa de Barros, from the swiss Support Center for Advanced Neuroimaging (SCAN, Inselspital, University of Bern). This meeting is part of a series of planned virtual meetings on the topics: preprocessing, quantification, and simulation software for MRS/MRSI.
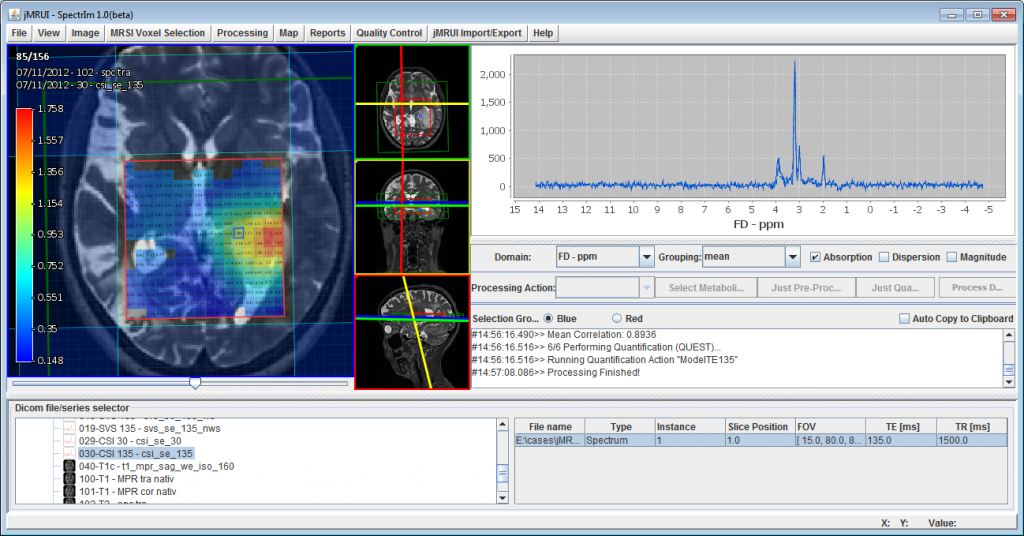
This virtual meeting is organized by the ISMRM Study Group on MR Spectroscopy and it will be held on Thursday 23 February at 07:00 PST, 10:00 EST, 16:00 CET. The meeting will start with a short introduction by Dr. Slotboom followed by a software demonstration by Nuno Pedrosa de Barros.
Note (*): This activity was restricted to members of the ISMRM MR Spectroscopy Study Group and required prior registration. To register, attendees had to go to the meeting registration page at ISMRM website and the login information was sent to registered attendees on Wednesday, 22 February 2017.
The virtual meeting video and the PowerPoint presentation are available for download at the ISMRM virtual meetings archive. Beware that a regular member login is required to access these files.

Post updated on 2019-01-08.