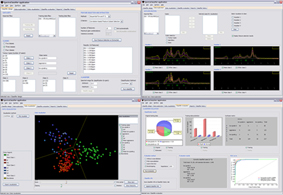
SpectraClassifier (SC) is a Java solution for designing and implementing Magnetic Resonance Spectroscopy (MRS)-based classifiers. The main goal of SC is to allow users with minimum background knowledge of multivariate statistics to perform a fully automated pattern recognition analysis. SC incorporates feature selection (greedy stepwise approach, either forward or backward), and feature extraction (PCA). Fisher Linear Discriminant Analysis is the method of choice for classification. Classifier evaluation is performed through various methods: display of the confusion matrix of the training and testing datasets; K-fold cross-validation, leave-one-out and bootstrapping as well as Receiver Operating Characteristic (ROC) curves.
SC is composed of the following modules: Classifier design, Data exploration, Data visualisation, Classifier evaluation, Reports, and Classifier history. It is able to read low resolution in-vivo MRS (single-voxel and multi-voxel) and high resolution tissue MRS (HRMAS) processed with existing tools (jMRUI, INTERPRET, 3DiCSI or TopSpin). In addition, to facilitate exchanging data between applications, a standard format capable of storing all the information needed for a dataset was developed.
SC is a user-friendly software designed to fulfil the needs of potential users in the MRS community.
Availability and requirements
| Plugin name: | SpectraClassifier (SC) |
| Project home page: | http://gabrmn.uab.cat/?q=sc |
| Operating system(s): | Platform independent |
| Programming language: | Java |
| Other requirements: | Java 1.6.0 or higher |
License
Available free of charge, subject to the signature of a disclaimer and user agreement text
Downloads and documentation
The SpectraClassifier 3.1.2 plugin and the related documentation can be downloaded from the project home page.
Support
Registered users should use the dedicated forum for all kind of support questions, or to report bugs and suggest new features and improvements.
If you have any problem downloading the software, please contact to:
Citing SpectraClassifier
We would appreciate from you to acknowledge SpectraClassifier in every instance where it has been used. It could be referenced by the following citation:
- SpectraClassifier 1.0: a user friendly, automated MRS-based classifier-development system. Sandra Ortega-Martorell, Ivan Olier, Margarida Julia-Sape and Carles Arus. BMC Bioinformatics 2010, 11:106. DOI:10.1186/1471-2105-11-106 (Published: 24 February 2010).